Metabolic profiling of plant, animal and human tissues and biofluids is an important technology in biology and medicine. The aim of these studies in humans is to use the profiles of metabolites, such as lipids, amino acids and sugars in the body, in a ‘systems biology’ approach, to understand changes that occur with events such as ageing, changing diets and the onset of diseases. Metabolic profiling is known as metabonomics: “the quantitative measurement of the multiparametric metabolic response of living systems to pathophysiological stimuli or genetic modification.” (Lindon et al, 2000). The alternate term metabolomics is also used interchangeably with metabonomics nowadays.
The health status of all organisms is the result of the interaction of genomic and environmental factors. Genomics can provide information on the contribution that our genes make to human health, disease states and the efficacy or otherwise of treatments for those disease states. However, genomics is blind to the host of environmental factors that affect humans such as: diet, taking medicines, exposure to environmental chemicals and critically, the status of our microbiome: the collection of microorganisms living in and on our bodies and particularly the gut microbiome. In addition, differences in a person’s genetic make-up only inform us about what might happen to that person in the future. With some notable exceptions, such as inborn errors of metabolism, there is a large degree of uncertainty and no direct correlation between many genomic differences and the physiological status of the individual.
On the other hand, metabolic profiling of human biofluids gives information in real time about the actual physiological status of an individual through analysis of their metabolites, and critically, these profiles reflect both genetic and environmental influences.
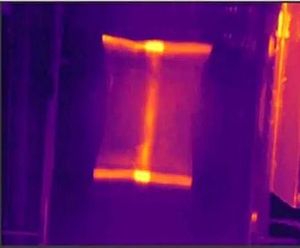
Metabolic profiles are measured using high-powered analytical technologies such as mass spectrometry (usually interfaced with a separation technology such as ultra performance liquid chromatography mass spectrometry (UPLC-MS)), or by nuclear magnetic resonance (NMR) spectroscopy (see Figure 1).
NMR spectroscopy
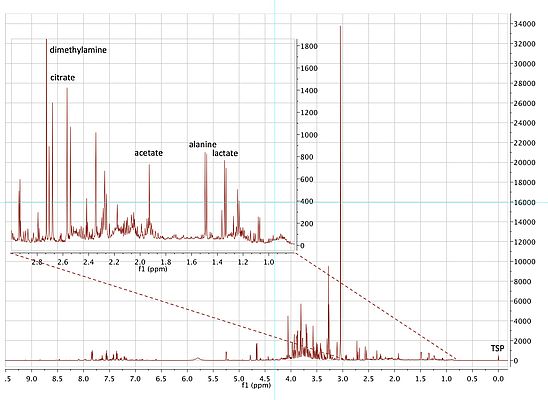
NMR spectra can be obtained from a variety of nuclei such as hydrogen-1, carbon-13 and nitrogen-15. Of those nuclei, hydrogen-1 is the most useful as it has high sensitivity and is present in all human metabolites. Although insensitive relative to mass spectrometry, hydrogen-1 NMR spectroscopy is fully quantitative and highly robust, as there is no contact between the sample and spectrometer. In addition NMR spectroscopy is the most powerful molecular structure elucidation technology for solution state samples. This is because the signals for each chemically distinct hydrogen occur at different characteristic chemical shift positions relative to an internal standard (Figure 2). In addition, hydrogens that are close in chemical bonds to other hydrogens or carbon-13 nuclei ‘couple’ with those nuclei and give exquisite information about their structural environment, thus facilitating metabolite identification (Dona et al, 2016). The representative hydrogen-1 NMR spectrum of the urine of a diabetic patient in Figure 2, shows the vast array of metabolite signals that can be observed.
Metabolic profiling or metabonomics/metabolomics is usually conducted in a diagnostic fashion. For instance, the metabolic profile of a person might be recorded before and after an intervention such as drug administration, and the effects of that drug on the metabolite profiles determined. This was the dominant paradigm for several decades (Lindon et al 2007). However a prognostic approach is also possible, where the metabolic profile is used to predict the outcome of a future intervention, that has not yet happened, which could be e.g. drug administration or just the passage of time.
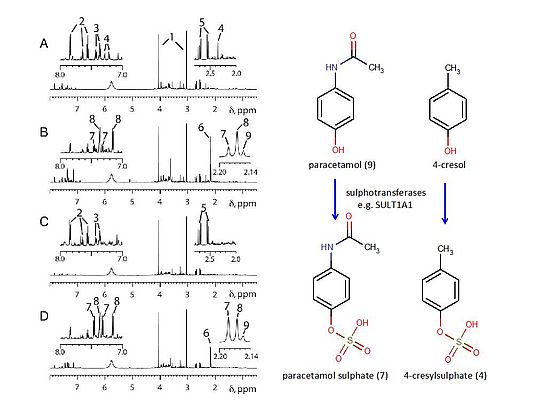
The first use of metabonomics to predict the effects of drug administration in humans (Clayton et al, 2009) demonstrated that high pre-dose levels of a gut microbiome derived metabolite were predictive of low post-dose levels of the sulphate metabolite of the drug paracetamol in a cohort of volunteers (See Figure 3).
Predictable drug effects
The pre-dose urine of volunteer 1 (Figure 3A) showed high levels of a microbial metabolite, (4), that was subsequently proven to be 4-cresol sulphate (Figure 3). After dosing with paracetamol, volunteer 1 excreted low levels of the paracetamol sulphate metabolite ((7), Figure 3B) relative to volunteer 2 (Figure 3D), who had no visible signals for 4-cresol sulphate (4) in their pre-dose urine spectrum (Figure 3C). This result was found to be generally applicable in the entire cohort of 100 human volunteers studied. The result is significant as: 1) it represents the first definitive human demonstration of pharmacometabonomics: the ability to predict drug effects based on pre-dose metabolite profiles; 2) the metabolism of a human drug is surprisingly shown to be partly under bacterial control, as 4-cresol sulphate is derived from the sulphation of 4-cresol, a gut microbiome metabolite and 3) as many drugs are metabolised by sulphation, the same gut microbiome drug metabolism influences may apply to them as well.
The development of prognostic metabolic profiling is significant for the advancement of personalised medicine, which is a key goal of 21st century medicine (Holmes et al Eds, 2016) and also significant for the pharmaceutical industry. Currently, most personalised medicine approaches are based on pharmacogenomics i.e. the use of genomics to predict drug effects in terms of efficacy, safety and metabolism, and these have had less success than originally hoped. It is anticipated that the use of pharmacometabonomics in concert with pharmacogenomics will improve our ability to predict drug effects and will help deliver the promise of personalised medicine; which is the selection of medicines for subgroups of patients so as to maximise drug efficacy and minimise toxicity. The use of pharmacometabonomics will also be important in the pharmaceutical industry where it could be used to help select participants in clinical trials so as to improve efficacy and reduce the occurrence of safety issues in those trials, as well as to provide a basis for patient selection post-launch so as to maximise patient benefits. The future is exciting and full of promise!
Author: Professor Jeremy R. Everett, Medway Metabonomics Research Group, University of Greenwich